Basic Usage Examples#
This notebook walks through representative data processing patterns to show what you can do with limulus.
import limulus
import pandas as pd
import polars as pl
iris_data = pd.read_csv('https://raw.githubusercontent.com/mwaskom/seaborn-data/master/iris.csv')
session = limulus.Session()
session.loads({"iris": iris_data})
Example 1: Common Syntax#
session.submit("""
data ratios;
set iris;
sepal_ratio = round(sepal_length / sepal_width, 0.01);
petal_area = round(petal_length * petal_width, 0.01);
keep species sepal_length sepal_width sepal_ratio petal_area;
run;
""")
print(pl.from_arrow(session["ratios"]).head(5))
shape: (5, 5)
┌─────────┬──────────────┬─────────────┬─────────────┬────────────┐
│ species ┆ sepal_length ┆ sepal_width ┆ sepal_ratio ┆ petal_area │
│ --- ┆ --- ┆ --- ┆ --- ┆ --- │
│ str ┆ f64 ┆ f64 ┆ f64 ┆ f64 │
╞═════════╪══════════════╪═════════════╪═════════════╪════════════╡
│ setosa ┆ 5.1 ┆ 3.5 ┆ 1.46 ┆ 0.28 │
│ setosa ┆ 4.9 ┆ 3.0 ┆ 1.63 ┆ 0.28 │
│ setosa ┆ 4.7 ┆ 3.2 ┆ 1.47 ┆ 0.26 │
│ setosa ┆ 4.6 ┆ 3.1 ┆ 1.48 ┆ 0.3 │
│ setosa ┆ 5.0 ┆ 3.6 ┆ 1.39 ┆ 0.28 │
└─────────┴──────────────┴─────────────┴─────────────┴────────────┘
Example 2: DO–END and Multiple Output Datasets#
session.submit("""
data large_petal small_petal;
set iris;
if petal_length > 2.0 then do ;
size_group = "large" ;
output large_petal;
end ;
else do ;
size_group = "small" ;
output small_petal;
end ;
run;
""")
# 2 ways convert to polars
print(pl.from_arrow(session["large_petal"]).head(3))
print(session.dataset("small_petal").to_polars().head(3))
shape: (3, 6)
┌──────────────┬─────────────┬──────────────┬─────────────┬────────────┬────────────┐
│ sepal_length ┆ sepal_width ┆ petal_length ┆ petal_width ┆ species ┆ size_group │
│ --- ┆ --- ┆ --- ┆ --- ┆ --- ┆ --- │
│ f64 ┆ f64 ┆ f64 ┆ f64 ┆ str ┆ str │
╞══════════════╪═════════════╪══════════════╪═════════════╪════════════╪════════════╡
│ 7.0 ┆ 3.2 ┆ 4.7 ┆ 1.4 ┆ versicolor ┆ large │
│ 6.4 ┆ 3.2 ┆ 4.5 ┆ 1.5 ┆ versicolor ┆ large │
│ 6.9 ┆ 3.1 ┆ 4.9 ┆ 1.5 ┆ versicolor ┆ large │
└──────────────┴─────────────┴──────────────┴─────────────┴────────────┴────────────┘
shape: (3, 6)
┌──────────────┬─────────────┬──────────────┬─────────────┬─────────┬────────────┐
│ sepal_length ┆ sepal_width ┆ petal_length ┆ petal_width ┆ species ┆ size_group │
│ --- ┆ --- ┆ --- ┆ --- ┆ --- ┆ --- │
│ f64 ┆ f64 ┆ f64 ┆ f64 ┆ str ┆ str │
╞══════════════╪═════════════╪══════════════╪═════════════╪═════════╪════════════╡
│ 5.1 ┆ 3.5 ┆ 1.4 ┆ 0.2 ┆ setosa ┆ small │
│ 4.9 ┆ 3.0 ┆ 1.4 ┆ 0.2 ┆ setosa ┆ small │
│ 4.7 ┆ 3.2 ┆ 1.3 ┆ 0.2 ┆ setosa ┆ small │
└──────────────┴─────────────┴──────────────┴─────────────┴─────────┴────────────┘
Example 3: BY Group Processing (RETAIN + FIRST. / LAST.)#
session.submit("""
data avg_petal;
set ratios;
by species;
retain area_sum 0 area_count 0;
if first.species then do;
area_sum = 0;
area_count = 0;
end;
area_sum + petal_area;
area_count + 1;
if last.species then do;
avg_petal_area = round(area_sum / area_count, 0.01);
output;
end;
keep species area_count avg_petal_area;
run;
""")
print(session.dataset("avg_petal").to_polars())
shape: (3, 3)
┌────────────┬────────────┬────────────────┐
│ species ┆ area_count ┆ avg_petal_area │
│ --- ┆ --- ┆ --- │
│ str ┆ f64 ┆ f64 │
╞════════════╪════════════╪════════════════╡
│ setosa ┆ 50.0 ┆ 0.37 │
│ versicolor ┆ 50.0 ┆ 5.72 │
│ virginica ┆ 50.0 ┆ 11.3 │
└────────────┴────────────┴────────────────┘
Example 4: Joining Data#
# Create a species lookup table
species_info = pl.DataFrame({
"species": ["setosa", "versicolor", "virginica"],
"category": ["small", "medium", "large"],
})
session.loads({"species_info": species_info})
session.submit("""
/* Vertical concatenation */
data iris_doubled ;
set iris(in=in1) iris ;
if in1 = 1 then tag = "first" ;
else tag = "second" ;
run ;
/* Horizontal join */
data iris_labeled;
merge iris(in=in1) species_info;
by species ;
if in1 ;
run;
""")
print("rows:", session["iris_doubled"].shape[0])
print(session.dataset("iris_labeled").to_polars().head(5))
rows: 300
shape: (5, 6)
┌──────────────┬─────────────┬──────────────┬─────────────┬─────────┬──────────┐
│ sepal_length ┆ sepal_width ┆ petal_length ┆ petal_width ┆ species ┆ category │
│ --- ┆ --- ┆ --- ┆ --- ┆ --- ┆ --- │
│ f64 ┆ f64 ┆ f64 ┆ f64 ┆ str ┆ str │
╞══════════════╪═════════════╪══════════════╪═════════════╪═════════╪══════════╡
│ 5.1 ┆ 3.5 ┆ 1.4 ┆ 0.2 ┆ setosa ┆ small │
│ 4.9 ┆ 3.0 ┆ 1.4 ┆ 0.2 ┆ setosa ┆ small │
│ 4.7 ┆ 3.2 ┆ 1.3 ┆ 0.2 ┆ setosa ┆ small │
│ 4.6 ┆ 3.1 ┆ 1.5 ┆ 0.2 ┆ setosa ┆ small │
│ 5.0 ┆ 3.6 ┆ 1.4 ┆ 0.2 ┆ setosa ┆ small │
└──────────────┴─────────────┴──────────────┴─────────────┴─────────┴──────────┘
Example 5: Integration with Python#
Here we combine Python processing before and after the Data Step.
As an example, we run a loop to compute sepal area by species and display scatter plots with matplotlib.
import matplotlib.pyplot as plt
for sp in ["setosa", "versicolor", "virginica"]:
session.submit(f"""
data sepal_{sp} ;
set iris ;
where species = "{sp}" ;
sepal_area = round(sepal_length * sepal_width, 0.01) ;
keep species sepal_area petal_length petal_width;
run ;
""")
out = session[f"sepal_{sp}"].to_pandas()
print(f"species={sp}: {len(out)} rows, mean sepal_area={out['sepal_area'].mean():.2f}")
plt.figure()
plt.scatter(out["petal_length"], out["petal_width"], alpha=0.5)
plt.yticks([0, 0.5, 1.0, 1.5, 2.0, 2.5])
plt.xticks([0, 2.5, 5.0, 7.5])
plt.show()
plt.close()
species=setosa: 50 rows, mean sepal_area=17.26
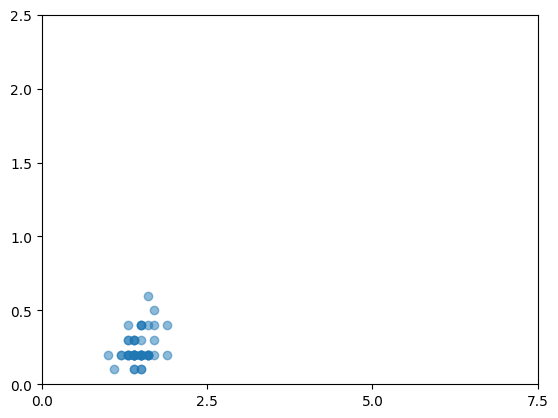
species=versicolor: 50 rows, mean sepal_area=16.53
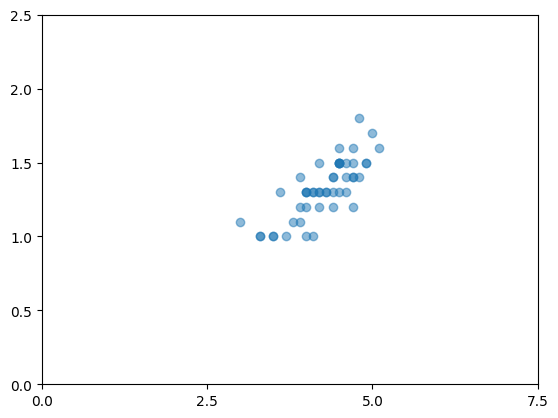
species=virginica: 50 rows, mean sepal_area=19.68
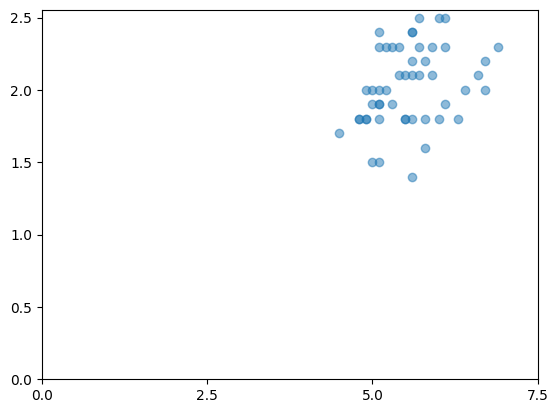
Example 6: Adding Label Metadata#
You can assign label information as Arrow metadata.
However, if additional processing is performed afterward, the metadata may currently be dropped unintentionally.For this reason, labels should be applied immediately before producing the final output.
session.submit("""
data iris_labels(label="Iris sample");
set iris(obs=3);
label sepal_length = "Sepal Length";
run;
""")
print(session.dataset("iris_labels").to_arrow().schema)
sepal_length: double
-- field metadata --
label: 'Sepal Length'
sepal_width: double
petal_length: double
petal_width: double
species: string
-- schema metadata --
memlabel: 'Iris sample'
Example 7: Column-Oriented API and Session Helpers#
It is intended to make it easier to express proc sytep operations and column-oriented processing needed for high-performance workloads.
This part is still under active development, and additional functionality will be added in upcoming updates. Column-oriented helpers on Session and DatasetView currently require exact column names and are case-sensitive.
After obtaining a dataset view with session.dataset(), you can use methods such as sort(), select() (for column selection and reordering), and astype() for type conversion.
(
session.dataset("iris")
.sort(by="species", out="iris_s")
.select(["species","sepal_length"])
.astype({"sepal_length": "float32"})
)
print(session.dataset("iris_s").to_polars().head(3))
shape: (3, 2)
┌─────────┬──────────────┐
│ species ┆ sepal_length │
│ --- ┆ --- │
│ str ┆ f32 │
╞═════════╪══════════════╡
│ setosa ┆ 5.1 │
│ setosa ┆ 4.9 │
│ setosa ┆ 4.7 │
└─────────┴──────────────┘
Data creation using SQL queries is supported through session.sql.
The SQL syntax follows the Polars SQL implementation.
session.sql(
"""
create table iris_summary as
select
species,
round(avg(sepal_length), 2) as avg_sepal_length,
round(avg(petal_length), 2) as avg_petal_length
from iris
group by species
"""
)
print(session.dataset("iris_summary").to_polars())
shape: (3, 3)
┌────────────┬──────────────────┬──────────────────┐
│ species ┆ avg_sepal_length ┆ avg_petal_length │
│ --- ┆ --- ┆ --- │
│ str ┆ f64 ┆ f64 │
╞════════════╪══════════════════╪══════════════════╡
│ versicolor ┆ 5.94 ┆ 4.26 │
│ setosa ┆ 5.01 ┆ 1.46 │
│ virginica ┆ 6.59 ┆ 5.55 │
└────────────┴──────────────────┴──────────────────┘
If you have code saved in a text file, you can load and execute it using session.include.
from pathlib import Path
from tempfile import TemporaryDirectory
with TemporaryDirectory() as tmpdir:
script = Path(tmpdir) / "include.txt"
script.write_text("""
data inc_sample;
set iris(obs=3);
run;
""", encoding="utf-8")
session.include(script)
print(session.dataset("inc_sample").to_polars())
shape: (3, 5)
┌──────────────┬─────────────┬──────────────┬─────────────┬─────────┐
│ sepal_length ┆ sepal_width ┆ petal_length ┆ petal_width ┆ species │
│ --- ┆ --- ┆ --- ┆ --- ┆ --- │
│ f64 ┆ f64 ┆ f64 ┆ f64 ┆ str │
╞══════════════╪═════════════╪══════════════╪═════════════╪═════════╡
│ 5.1 ┆ 3.5 ┆ 1.4 ┆ 0.2 ┆ setosa │
│ 4.9 ┆ 3.0 ┆ 1.4 ┆ 0.2 ┆ setosa │
│ 4.7 ┆ 3.2 ┆ 1.3 ┆ 0.2 ┆ setosa │
└──────────────┴─────────────┴──────────────┴─────────────┴─────────┘